Free Release of BacMicroSim Version 1.0
In Silico Fermentation has been developing freely downloadable MATLAB applications that streamline the utilization of microbial metabolic modeling methods. To date, we have released four research oriented apps (MediumFBA, InSilicoKO, STBRsim, SynComSim), and one educationally focused app (STBRtutorial).
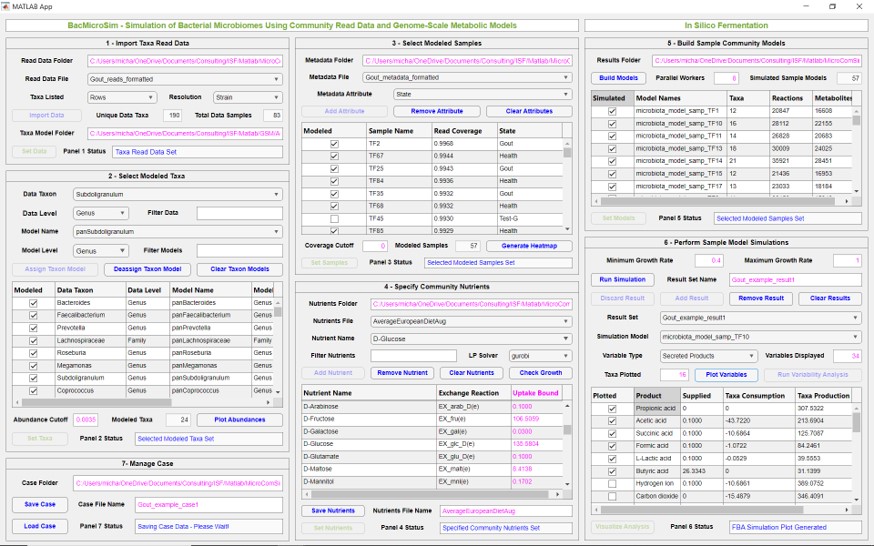
Now we are pleased to announce the release of BacMicroSim, a free MATLAB application that enables rapid development, simulation and analysis of bacterial community models derived from taxonomic read data. Given genome-scale metabolic reconstructions of the relevant taxa, the GUI tool combines abundance data and nutrient uptake constraints to form sample-specific community models that can be subjected to flux balance analysis (FBA) and flux variability analysis (FVA) to predict nutrient uptake, product secretion, and metabolite crossfeeding rates. All predictions are resolved on both a community and taxa-by-taxa basis.
The workflow allows importing of Excel read and metadata files, definition of the taxa and samples to be modeled, supplied nutrients and their uptake bounds, automated building of new sample models, selection of sample models to be simulated, and definition of metabolite fluxes for FVA. Community simulations can be performed for different combinations of user-defined parameters and stored for plotting and comparison using the included visualization App.
More information about BacMicroSim and the freely downloadable application are available here.