Training Course on Genome-Scale Metabolic Reconstructions and Flux Balance Analysis
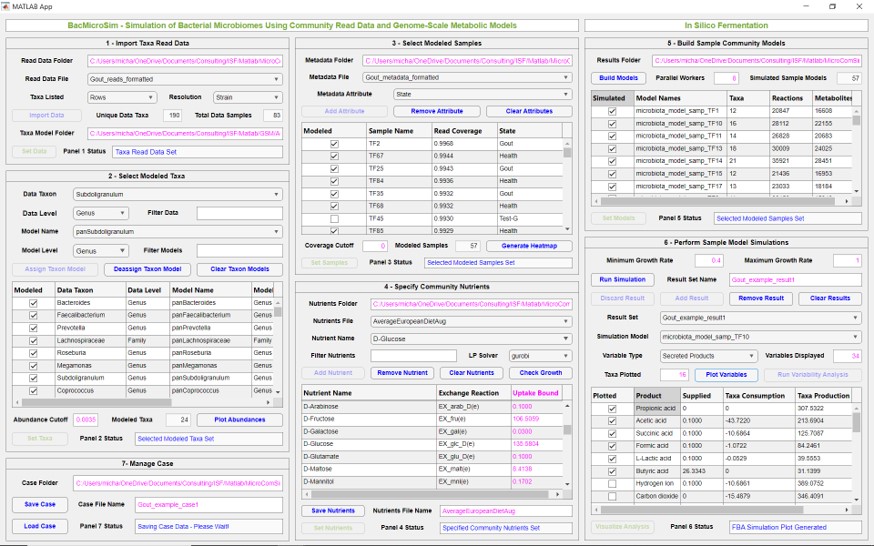
In Silico Fermentation is pleased to announce our first training course focused on genome-scale metabolic reconstructions and their use for constraint-based metabolic analysis. The training focuses on developing a functional knowledge of metabolic reconstructions and more in-depth understanding of flux balance analysis (FBA) and its most important variants. Key concepts are demonstrated through computational exercises […]