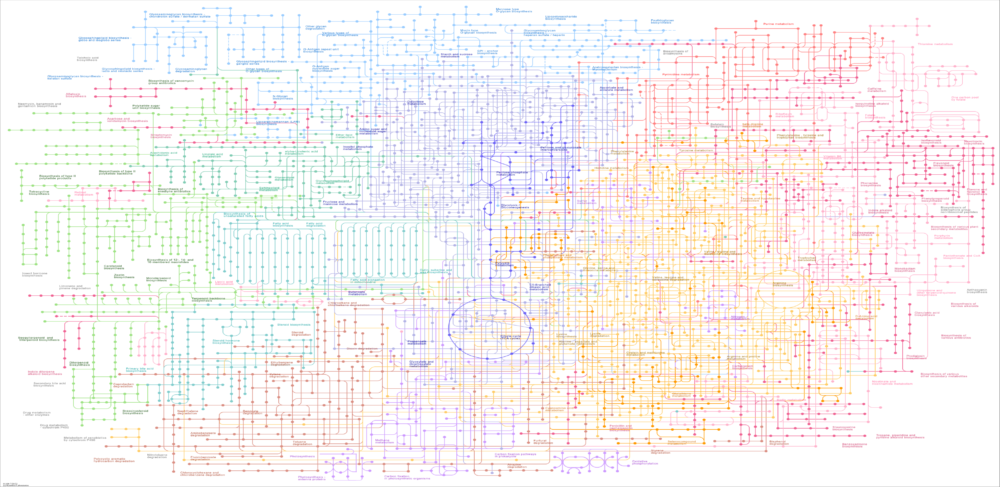
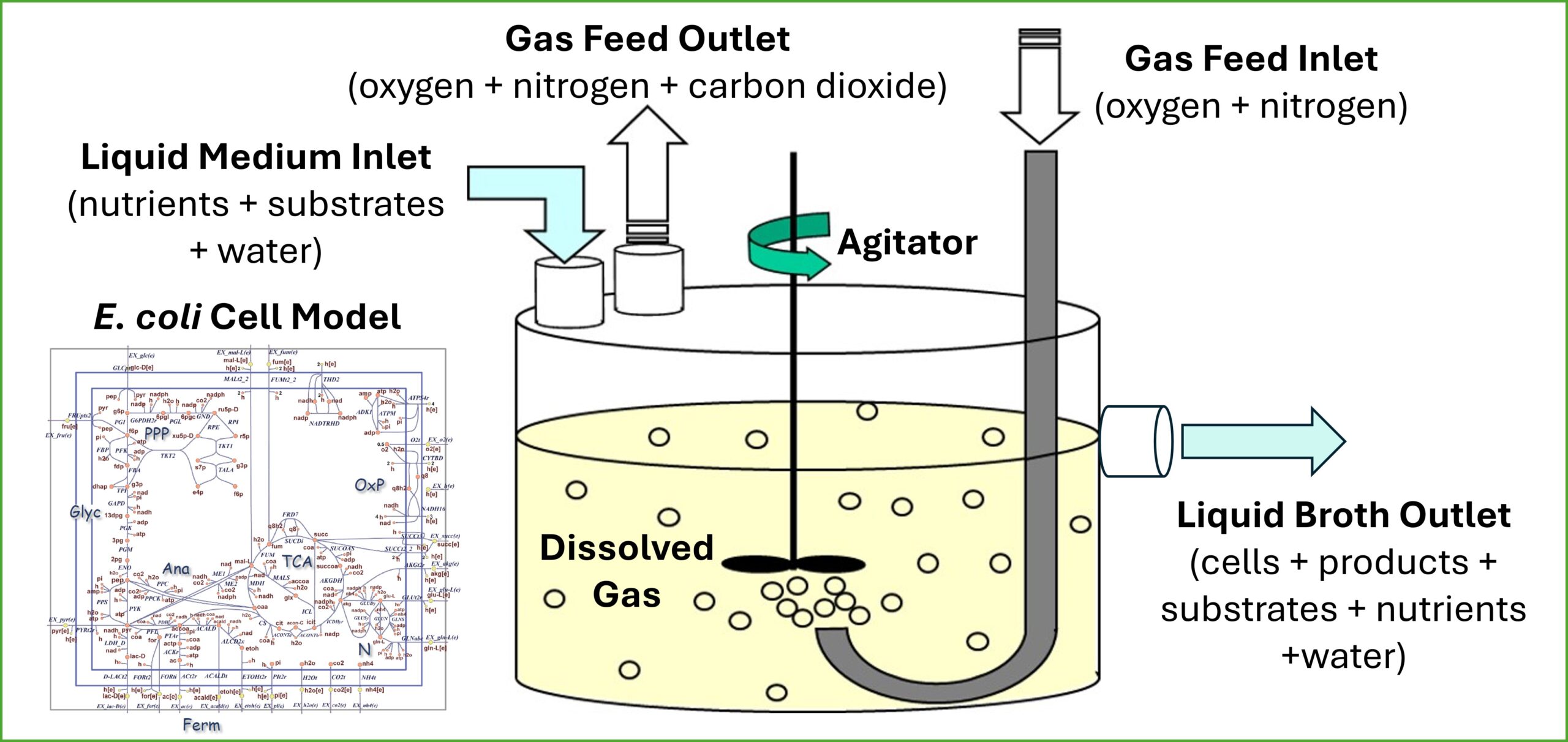
While genome-scale metabolic reconstructions are routinely used as a computational platform for microbial strain analysis and design, their incorporation within process-oriented dynamic models is less common. Using dynamic extensions of flux balance analysis, metabolic reconstructions can be embedded within dynamic descriptions of the extracellular environment to develop simulation models of stirred tank bioreactors for predicting and enhancing fermentation performance. To effectively utilize these simulation models, users must have adequate understanding of the bioreactor model structure, the adjustable model parameters, and the computational methods used for model solution. Without such knowledge, users often struggle to generate successful simulation runs, interpret individual simulation results, and to understand steady-state and dynamic trends as model parameters are changed.
The goal of this training course is to provide participants with the functional knowledge and hands-on experience needed to effectively utilize STBRsim, our MATLAB application for performing stirred tank bioreactor simulations based on genome-scale metabolic reconstructions. The training provides a conceptual understanding of the extracellular model component with an emphasis on the model structure and adjustable parameters. This knowledge allows participants to understand the app workflow and the process-relevant parameters that must be specified. The underlying simulation methods are explained at a level appropriate to configure and troubleshoot the app. Building upon this foundation, STBRsim is applied to three bioreactor simulation problems that differ with respect to the metabolic reconstruction, the process objective, and the bioreactor operating mode. The final portion of the course allows participants to configure and solve their own bioreactor simulation problem, with assistance from the instructor as needed. Participants complete the course with a fully configured app and the hand-on experience necessary to pursue bioreactor simulations in their future process development efforts..
Course Topics
1. Formulation of stirred-tank bioreactor models
2. Dynamic simulation of bioreactor models
3. Specification of adjustable model parameters
4. Utilizing STBRsim for stirred-tank bioreactor simulation
Course participants are expected to have completed the training course Genome-Scale Metabolic Reconstruction and Flux Balance Analysis. Also recommended is practical experience with:
1. Gas-liquid mass transfer
2. Ordinary differential equations
3. Unstructured bioreactor modeling
Please contact me by email if you would like to learn more about this training course.