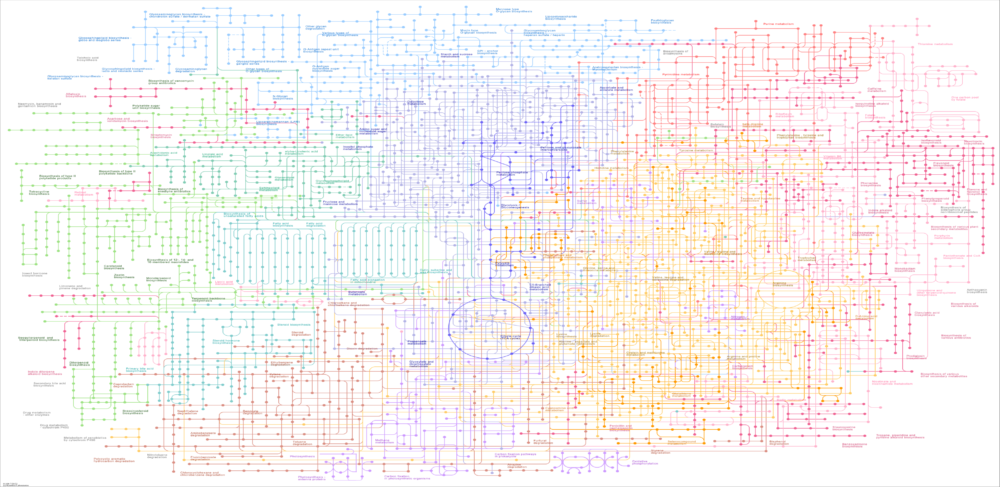
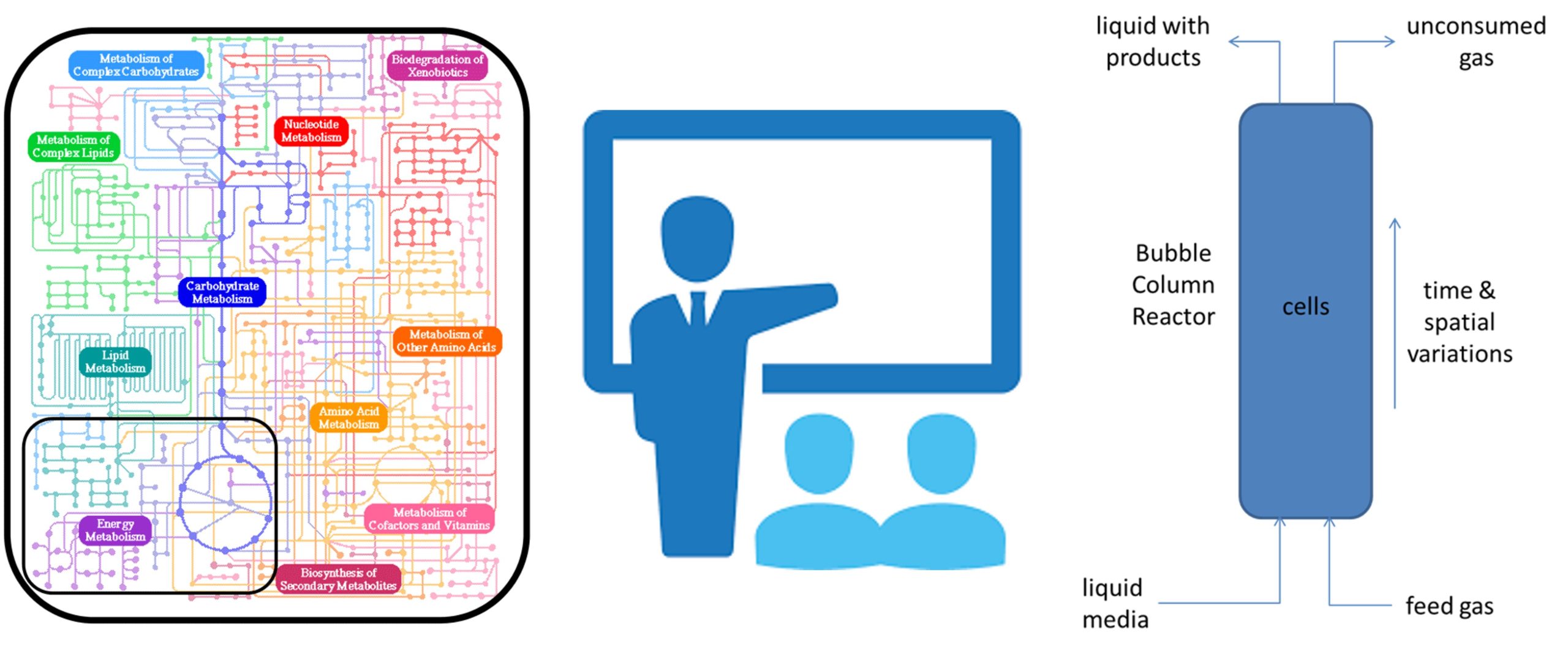
Genome-scale metabolic reconstructions and associated constraint-based modeling methods have emerged as powerful tools for microbial system analysis and design. Metabolic modeling is often used to compliment in vitro experiments and in vivo data collection by generating predictions that are difficult or impossible to generate by other means. For the most part, the use of these in silico tools has been limited to academic researchers or previously trained industrial practitioners. As a result, companies often lack the in-house expertise necessary to incorporate metabolic modeling into their research and development efforts.
To bridge this knowledge gap, In Silico Fermentation is offering focused training courses aimed at providing practitioners with the essential background and hands-on experience required to effectively utilize metabolic modeling. The courses deliver functional knowledge about genome-scale metabolic reconstructions, flux balance analysis (FBA) and other constraint-based modeling methods, in siilco metabolic engineering, bioreactor dynamic simulation, and microbial community modeling. For each topic, the basic concepts are presented and then simple computational examples are completed to demonstrate their implementation. A major focus of each course is the extensive use of hands-on exercises using metabolic reconstructions chosen by the participants. The courses are designed for on-line delivery in time blocks defined by the participants to maximize work flexibility and information retention.
The following training courses have been developed and are available for immediate delivery:
- Genome-scale metabolic reconstructions and flux balance analysis
- Bioreactor modeling using genome-scale metabolic reconstructions
Additional courses are under development, and customized courses can be offered upon request.
Please contact me by email if you would like to learn more about these metabolic modeling training courses.